Autism Spectrum Disorders
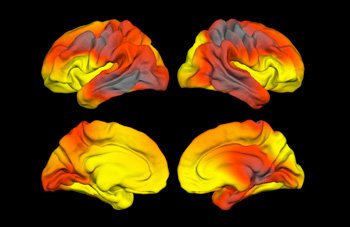 In the top row are lateral views and in the bottom row are mesial views showing the association of MECP2 SNP-rs2239464 with cortical surface area in males. The map shows the distribution of nominal -log p-values (sign indicating direction of effect per copy of minor allele) across the reconstructed cortical surface. STSI researchers did not detect significant effects in the corresponding map for females.
In the top row are lateral views and in the bottom row are mesial views showing the association of MECP2 SNP-rs2239464 with cortical surface area in males. The map shows the distribution of nominal -log p-values (sign indicating direction of effect per copy of minor allele) across the reconstructed cortical surface. STSI researchers did not detect significant effects in the corresponding map for females.
Several genes have been linked to autism spectrum disorders (ASD), with one of the most intriguing being MeCP2 (methyl-CpG-binding protein 2), a very complex and multifunctional nuclear protein whose mechanism of action is not yet fully understood.
MeCP2 first attracted scientific attention just over a decade ago when a Baylor College of Medicine investigator discovered that mutations in this gene cause Rett’s Syndrome (RTT), a member of the ASD category along with classic autism and Asperger Syndrome. Subsequently, altered MeCP2 expression was detected in the cells of some individuals with autism.
The gene’s location on the X chromosome may help explain autism’s higher prevalence in males, who have only one X chromosome while females have a pair. Since males with the gene mutation do not have a “back up” X chromosome, the impact of the variation may be greater for them than it is for females who have the mutation.
In Dec. 2009, Scripps Translational Science Institute (STSI)’s investigators were part of the international team of investigators who revealed that common sequence variations in MeCP2 show association with regional measures of brain cortical surface area, predominately in frontal and anterior temporal cortices. They reported their findings in the Proceedings of National Academy of Sciences“>Proceedings of the National Academy of Sciences in a paper whose lead author was Alexander Joyner, Ph.D., a former STSI postdoctoral researcher.
With DNA extracted from cells of 73 two- to six-year-old patients with autism at UC San Diego’s Autism Center of Excellence and Healthy Infant Development Laboratory, STSI scientists used next-generation sequencing technology to assess variation in the 164 kilo base pair (kb) region including MeCP2 and regulatory elements. They will search the DNA sequences of people without ASD to determine whether healthy individuals’ genomes possess any of the novel variants and other single nucleotide polymorphisms (SNPs) and insertions/deletions that they’ve detected in the DNA sequences of the 73 children with autism. The control group’s DNA was provided by the 1,000 Genomes Project.
With access to brain images and the medical histories of the 73 children, STSI scientists hope to determine whether the strongest genetic mutations, or even those that are relatively common in the population, are associated with the abnormal neurodevelopmental features of autism.
